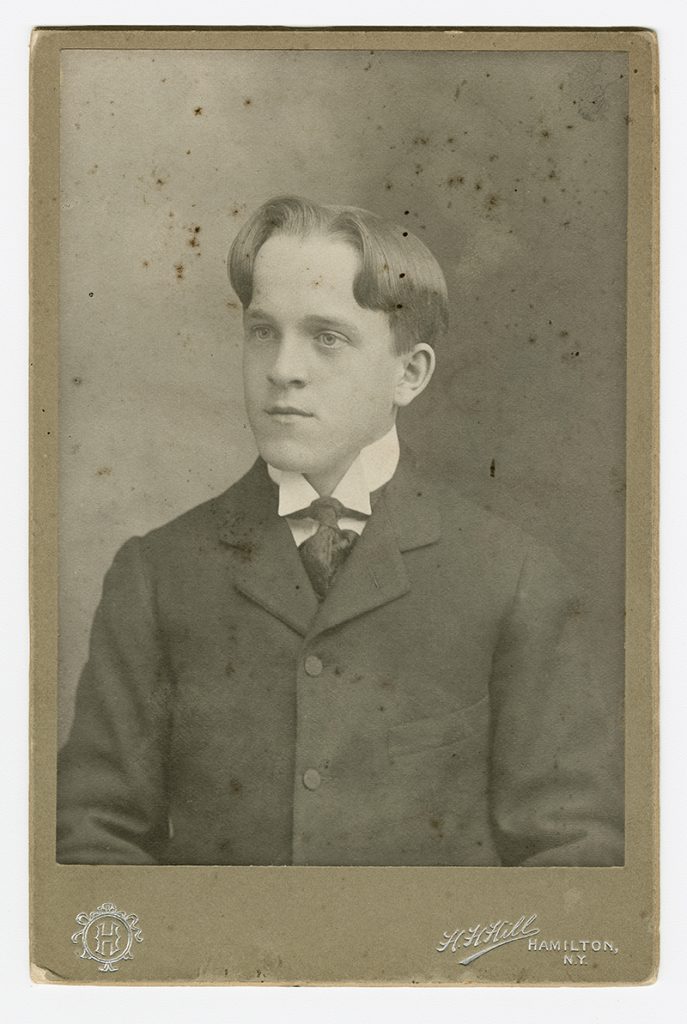
Oswald Avery, Class of 1900, made a discovery that revolutionized science. Now, Colgate biologists are following in his footsteps to make their own genetic revelations.
Oswald Avery, Class of 1900, was a man who collected nicknames. During his years at fin de siècle Colgate, fellow students called him “Babe” due to his slight stature — he rarely weighed more than 100 pounds — and boyish features. As an immunology researcher at Rockefeller Institute for Medical Research in New York, colleagues called him “Fess,” short for “Professor,” for his enthusiastic expositions on science, “always spicing his performance, with mimicry, pithy remarks, picturesque analogies, and verbal pyrotechnics,” in the words of one. One name Avery never collected, however, is “Nobel Prize Winner.” And yet, he made arguably the most significant discovery in 20th-century biology: the realization that a chemical called DNA carried the genetic information to control the development of all organisms.
Avery published that bombshell in the obscure Journal of Experimental Medicine in 1944 — nearly a decade before James Watson and Francis Crick’s breakthrough of the DNA double-helix assured their own place in Nobel history. In part due to Avery’s self-deprecating manner, which made him cautious of trumpeting his finding too loudly, colleagues ignored and disputed his discoveries for years. But he was under no illusions about the importance of his breakthrough. “If we are right,” he wrote in a letter to his younger brother Roy in 1943, “nucleic acids are not merely structurally important, but functionally active substances in determining the biochemical activities and specific characteristics of the cells — and that, by means of a known chemical substance, it is possible to induce predictable and hereditary changes in cells.” In the 75 years since his discovery, that’s exactly what scientists have done.
Avery was born in Halifax, Nova Scotia, on Oct. 21, 1877. He was the son of a Baptist minister, who brought him to New York City at age 10. Precocious as a child, Avery played the cornet on Sundays to attract parishioners to his father’s Lower East Side church. He attended Colgate Academy, and then Colgate University, where he led the University band with his trumpet and won competitions for public speaking. Despite excellent grades in music and the humanities, however, he only took the required science classes. So it surprised everyone when, upon graduation, he decided to pursue a degree at Columbia Medical School. Working with patients as a doctor in clinical practice, Avery felt frustrated at how little he could do for those with chronic diseases, and he decided to turn to research instead — first at Hoagland Laboratory in Brooklyn and then at the Rockefeller Institute in 1913.
He made arguably the most significant discovery in 20th-century biology: the realization that a chemical called DNA carried the genetic information to control the development of all organisms.
Avery didn’t set out to discover DNA. His main interest was finding a cure for pneumonia, which at the time was killing thousands of people every year. In a small, sixth-floor laboratory at the Rockefeller Institute, he pursued the bacteria that caused it, Streptococcus pneumoniae, for the better part of 35 years. Colleagues reported on two sides to Avery’s personality. On the one hand, there was the gregarious “Fess,” who was always smiling and happy to talk with his labmates. On the other was a quiet, almost brooding scientist, who eschewed big gatherings and rarely traveled, other than yearly trips to Maine. This Avery spent hours in the lab behind round spectacles, meticulously planning experiments to isolate and attack his nemesis.
Pneumonia comes in two forms: smooth (S), which is virulent, and rough (R), which is not. In 1923, Avery was the first to discover the difference between the two: a polysaccharide sugar coating on the S strain that protected the pathogen from being killed by a host’s immune system. An experiment by a British biologist in 1928 had shown that, by injecting a mouse with a non-virulent R strain along with a type S strain that had been previously killed (and was therefore not pathogenic), he could create a live S strain bacteria, causing the mouse to die within days. Such a change was unheard of. Clearly some “transforming principle” inside the dead bacteria was turning the live bacteria into a killer, but what was it?
Scientists had long understood the concept of genes, which determine how traits of organisms are inherited, but in the 1940s, they considered them to be an abstract concept, with little understanding of what chemicals made them up within a cell. If they had to guess, most scientists would have assumed they were made of proteins in the nucleus. Along with two colleagues, Colin McLeod and Maclyn McCarty, Avery now set out to change that, tackling his goal in a patient, meticulous manner.
The researchers started with a culture made from beef hearts, in which they grew S strain pneumonia bacteria. They then killed the cells by breaking down their membranes, and washed off the sugar capsule with multiple applications of saline. Precipitating what was left in an alcohol solution, they removed most of the proteins with chloroform to create a small amount of viscous precipitate, which they discovered was mostly sodium salt of deoxyribonucleic acid — DNA. By injecting this into mice along with the non-virulent R-type bacteria, they achieved the same deadly effects as if they had injected the S strain bacteria. In further experiments, they used enzymes to digest any remaining protein and found the experiment still worked; but when they used enzymes to remove the DNA, it failed.
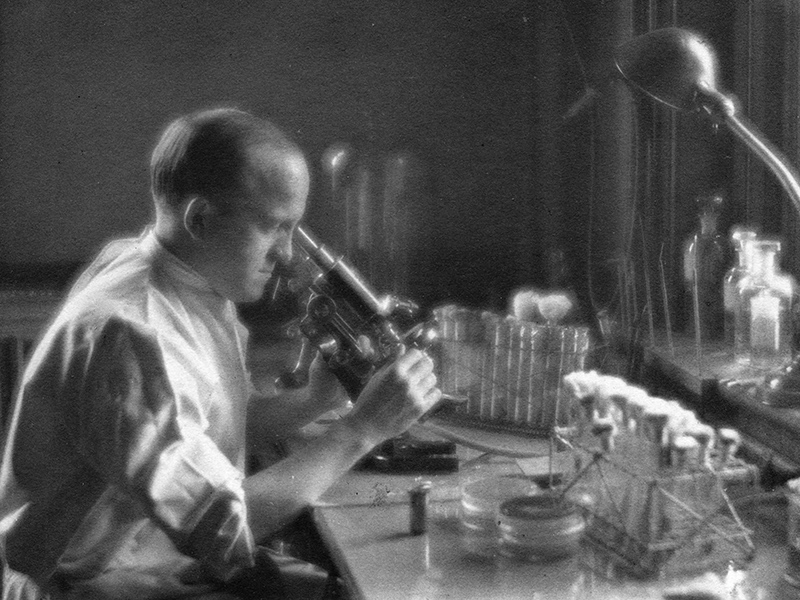

Avery and his colleagues wrote their paper with a cautious tone, saying only that the nucleic acids “must be regarded as possessing biological specificity the chemical basis of which is as yet undetermined.” Others were much more excited. When Avery won the New York Academy of Medicine gold medal in 1944 for his pneumonia research, the award committee noted the paper’s “very far-reaching implications.” A leading biochemist immediately grasped the possibilities of injecting DNA into cells to change their properties, saying “the importance of these observations can scarcely be overestimated.”
The bulk of the scientific community, however, failed to embrace the significance, fixated on the idea that the solution still contained trace amounts of protein, which they figured must really be behind the change. DNA, they thought at the time, consisted of only a monotonous sequence of bases, which couldn’t possibly account for the incredible genetic variability of life. Over the next few years, however, scientists began to replicate Avery’s experiment with similar results, and most came around to the idea that DNA was indeed the basis for genes in the cell. By the time Watson and Crick demonstrated that the double-helix structure of DNA could provide the needed variability along with a means for replication, most scientists had realized DNA’s singular importance inside the cell.
Since Avery’s time, we have been learning more and more about the molecular mechanisms operating within the brain and body. To use this information as a tool to look at things like behavior is really exciting.
— Krista Ingram, associate professor and biology department chair
That recognition was too late for Avery, who retired in 1948 and moved in with his brother Roy in Nashville. He died of liver cancer in 1955, at age 77. Chemist Arne Tiselius, winner of the Nobel Prize in 1948, called it “lamentable” that Avery never received the top award in his field. “Had he not died when he did, I think he most certainly would have gotten it,” he said. Recognition has since come, if slowly. In 1983, Esquire named Avery as one of the 50 “who made a difference” in the preceding half-century, alongside such 20th-century pioneers as Muhammad Ali, Walt Disney, Martin Luther King Jr., and Dr. Jonas Salk. Since then, the fields of microbiology and genetics have grown ever-more complex, with new tools and techniques to manipulate DNA to cure diseases, regenerate cells, and understand human and animal development like never before. But at the basis is Avery’s discovery in his lab 75 years ago. As the work of four modern-day researchers at Colgate shows, his legacy lives on, at his alma mater and beyond.
About Time
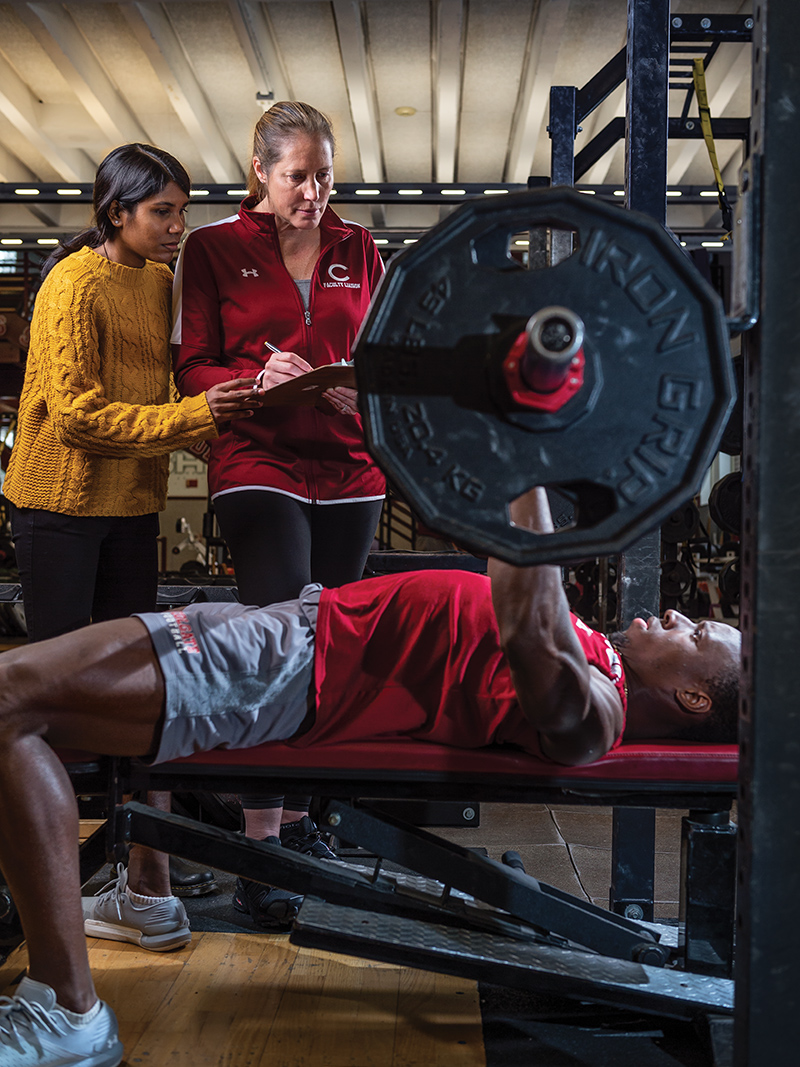

Everyone suspects they are either a morning person or a night owl. According to research by Associate Professor and Biology Chair Krista Ingram, that distinction might be genetic — hardwired into our DNA. Ingram has long been interested in the effects of genes on behavior, a much more difficult phenomenon to investigate than physical characteristics. “If you are looking at physical traits like eye color or freckles, you can often explain much of the variance in the trait due to heredity,” she says. “Looking at behavior and genetics wasn’t even possible until I was in graduate school, when microarrays came out and we could look at a suite of 8,000 genes at once.”
She began her career looking at behavior in insects, investigating something called the “foraging gene,” which seemed to control whether fruit fly larvae moved around a lot or stayed in place. In ants, this gene is associated with the daily timing of foraging outside the nest. Ingram, who is a morning person, was spurred to examine human circadian rhythms when her husband, a psychologist and a night owl — wondered if there were any genetic basis to those preferences. “I said, ‘Of course, they’ve looked into this,’ but I found out they hadn’t. I couldn’t believe it,” she says.
Time-of-day preference, or chronotype, comes down to a small number of genes, including a core group of eight that biologists call the clock genes. These genes transcribe proteins in a part of the brain called the suprachiasmatic nuclei, and then break them down on a 24-hour loop, regulating a daily cycle. Only, that’s not exactly true, Ingram found when she began researching the process. “What happens is, it’s not exactly 24 hours — you can be a little slow or a little fast — and that, in part, is what influences your behavior and influences whether you are a morning person or a night person,” Ingram says.
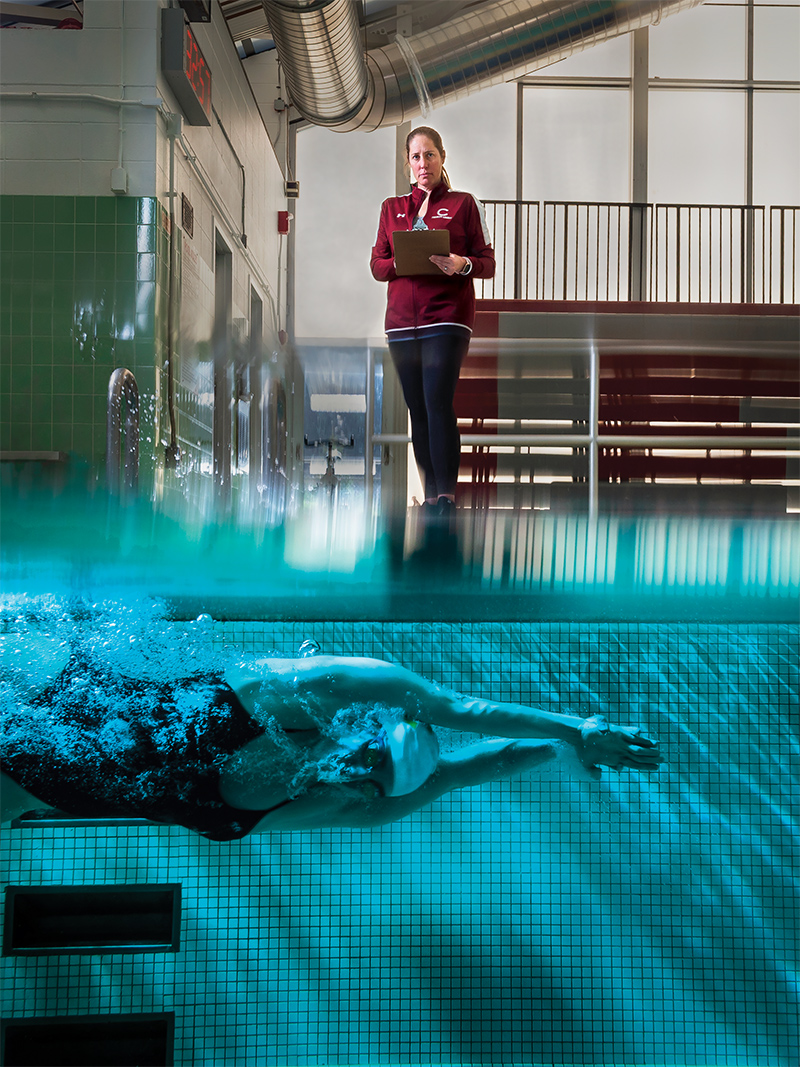

By examining gene transcription in hair follicles, Ingram has been able to tell where a person’s chronotype falls on the circadian spectrum. Only about 10–15% of people on either side are true morning or night people, she says, while about 70–80% fall somewhere in between. In looking at what causes this difference, Ingram has traced it to a genetic phenomenon called single nucleotide polymorphism (SNP, pronounced “snip”), in which a single nucleotide on a clock gene is switched — for example, a cytosine (C) instead of adenine (A). “If you have three or more of those SNPs, you have a higher probability of being a morning person,” she says.
That difference can have real-world repercussions. Ingram’s research has found that having a chronotype different from one’s social group — for example, a night person forced to wake up early for work or school — is associated with a higher prevalence of anxiety and depression. In addition, she has found that people are more likely to engage in risky or unethical behavior during the opposite time of day they prefer. “If you think about judges or investment bankers who are making important decisions involving large financial consequences, it may make a big difference whether they are at their circadian peak or not.”
Recently, Ingram has begun examining student-athletes at Colgate, to see how much chronotype makes a difference on their performance. In tests with the swim team, she has found that swimmers competing off their circadian peak can post times a few hundredths of a second slower than they do when swimming at their preferred time of day. In a sport where fractions of a second make a big difference, such results argue for the importance of understanding chronotype on all of our performance. “We all need to understand that there are times of day when we will have to work harder to achieve the same results,” Ingram says. By becoming more aware of our genetically preferred time of day, she says, we can better plan important tasks, and put in more effort when we know we are working off our peak.
A Dog’s Life
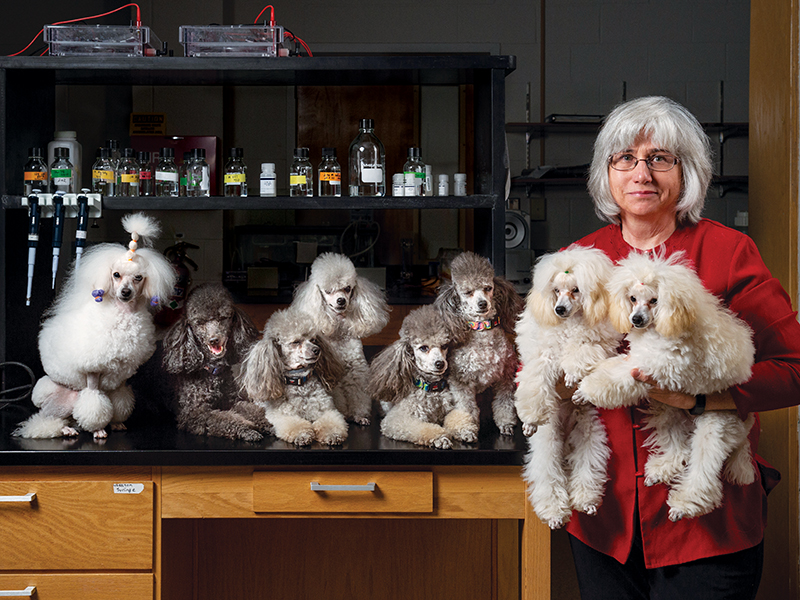
Biology professor Barbara Hoopes didn’t expect her scientific career to lead her into the ring at the Westminster Kennel Club dog show, running alongside her toy poodle Tommy in a master agility competition. But for Hoopes, the experience combines her two great passions — dogs and genetics. Hoopes showed poodles at dog shows when she was young, enjoying grooming them and teaching them tricks. That pastime fell by the wayside in college as she traded dogs for simpler species such as bacteria and yeast, and seeing what she could trick them into doing when their DNA was changed.
“I love being able to manipulate the genome and see what effect it has,” she says. After earning her tenure as a professor, she began showing dogs again — and thought it might be interesting to examine their genetics as well. She began working with the National Institutes of Health (NIH) to study how genes affect dog size, and in the midst of collecting samples at a poodle national dog show, she became enamored with a little silver toy poodle performing agility challenges. “Now I have 11 of them,” says Hoopes, who in 2015 was declared an American Kennel Club breeder of merit. In addition to Tommy, who placed third at the Westminster show, she now competes with dogs of her own breeding. Six of the 11 pups she has bred since 2014 are champions or agility champions.
What fascinates Hoopes about poodles is the incredible diversity of size in one breed, which ranges from 8 inches at the shoulder for a toy poodle all the way up to 25 inches for a standard poodle. She knew there had to be something interesting happening genetically to allow for that variation. “I thought to myself, how are these small dogs even alive?” she remembers. “What other compensating mutations are allowing them to survive?” While she was leading the NIH study group, Hoopes and other NIH researchers found that a small number of genes could provide for this variety. In particular, Hoopes has focused on the effect on a single mutation in a gene called growth factor 1 receptor (GF1R). “That gene and five others can explain 80% of the variation in breed-specific height,” she says.
Avery made an important contribution to science with a really elegant experiment. We let students know they can all play a part in contributing to science.
— Barbara Hoopes, biology professor
Insights like that can help breeders by figuring out in advance the range in possibilities of size in a litter of puppies from two parents. Toy poodles, for instance, can range between 3 to 6 ounces at birth. “The really big puppies are difficult for a toy mother to naturally give birth to, while the really small ones are hard to keep alive, and we have evidence GF1R is important to this variation,” Hoopes says. “So we are hoping our work will actually help breeders to know what to expect when they breed two dogs.” In her classes at Colgate, Hoopes helps students design their own molecular experiments with dogs, focusing on the effects of a particular gene. “There are all these questions no one has gotten an answer to yet,” Hoopes says. “The techniques are relatively straightforward, and undergrads can get some results and learn about doing science.”
Hoopes is now serving as a breeder representative for a toy poodle genetic diversity study sponsored by University of California-Davis. Having an understanding of genetics, she says, gives her a leg up in breeding her own dogs. But the results she is garnering from her research are not limited to canines. While dog size might be affected by a couple dozen genes overall, size in humans is controlled by hundreds. Nevertheless, she says, “if we can understand more about what these genes are doing in dogs, then it might help us understand more about what they are doing in humans and other species.”
Little Things That Count
Since Avery’s discovery of DNA, scientists have had a straightforward view of genetics: Organisms inherit genes composed of DNA from their parents, and that DNA is transcribed into messenger RNA, which creates proteins that build and influence cells. Thus, RNA was thought to be just a messenger that helps make the more important proteins. Two decades ago, however, a microscopic worm called C. elegans complicated that model, says Associate Professor of Biology Priscilla Van Wynsberghe. Scientists discovered a class of small RNA called microRNA that are essential for cellular function. “They can bind to messenger RNA to inhibit gene expression,” Van Wynsberghe says. “Small changes in gene expression can have devastating consequences on an organism.”
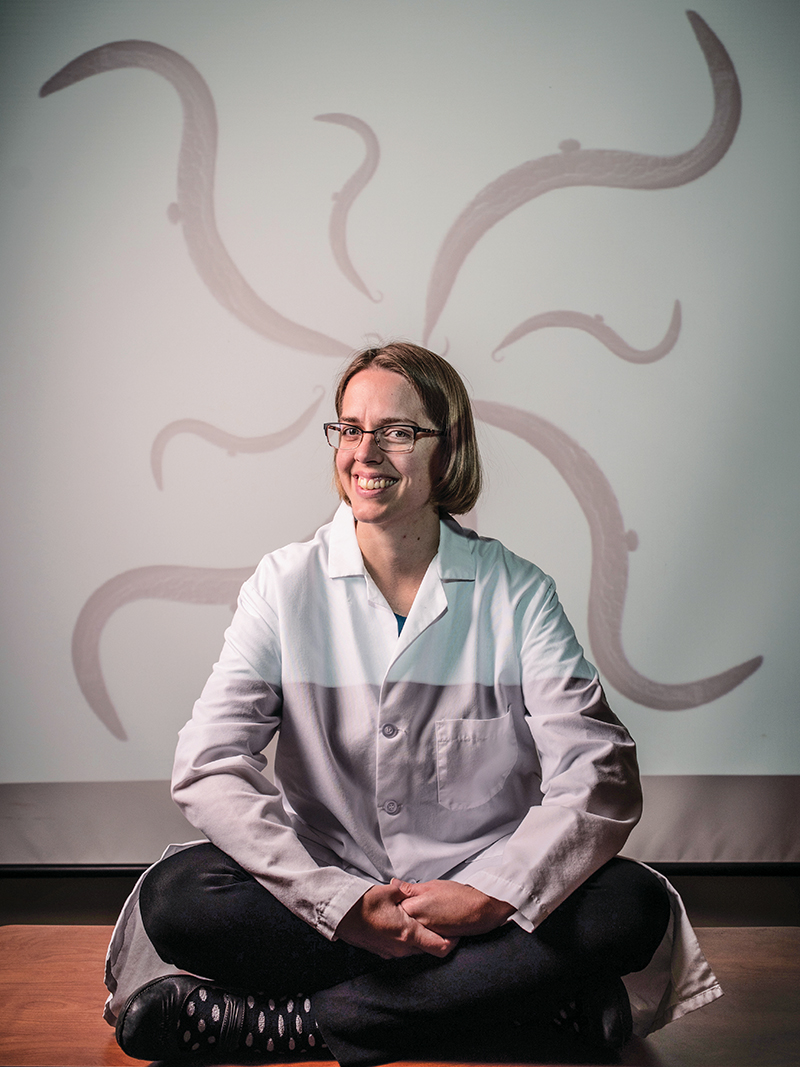

When one particular microRNA gene called let-7 is misregulated, for example, it can cause the worm to explode. “Its gonads and intestine literally explode out of its vulva,” says Van Wynsberghe. She conducts experiments on the species using CRISPR, a relatively fast and easy way to edit genes, which uses an enzyme to snip DNA to create a deletion or insert a new piece of DNA in its place. Understanding the role microRNAs play in the worm’s development can help scientists understand its role in humans because 35% of C. elegans genes have a human equivalent. “In humans, a misregulation of the let-7 microRNA has been associated with multiple cancers, including lung and colon cancer,” Van Wynsberghe says.
At Colgate, Van Wynsberghe works with students to design experiments using CRISPR and other molecular genetic techniques to manipulate microRNAs and other genes, thereby examining the effects on the worm’s development in real time. In addition, she has explored the effects of other components in the cell’s nucleus on worm development. For most of the history of genetics, scientists have believed that while DNA is passed on to subsequent generations, any modifications to cells during a parent’s lifetime dies with them. “We now know that some modifications to DNA, which do not involve DNA sequence changes, can be inherited as well,” Van Wynsberghe says. She and her students are studying this phenomenon called epigenetics. For example, they have looked at the effects of DES (diethylstilbestrol), a drug once given to women in the 1950s and 1960s to counteract pregnancy complications, which has since been associated with an increase in cancer. Daughters of women who took the drug while pregnant have been shown to be at higher risk for cancer, and their own daughters have shown to be at higher risk as well. In her experiments, Van Wynsberghe is studying how many generations of C. elegans show fertility and other defects after initial exposure to DES, as well as the underlying changes in DNA modification and gene expression that cause these effects. Her team has also been conducting experiments with other chemicals, such as plasticizers (compounds added to plastics to make them more pliable), and finding that an exposure to C. elegans can cause physical changes to several future generations. “The fact that these DNA modifications can be inherited and influenced by environmental signals,” she says, “is in some ways terrifying, but also fascinating.”
Miracle Grow
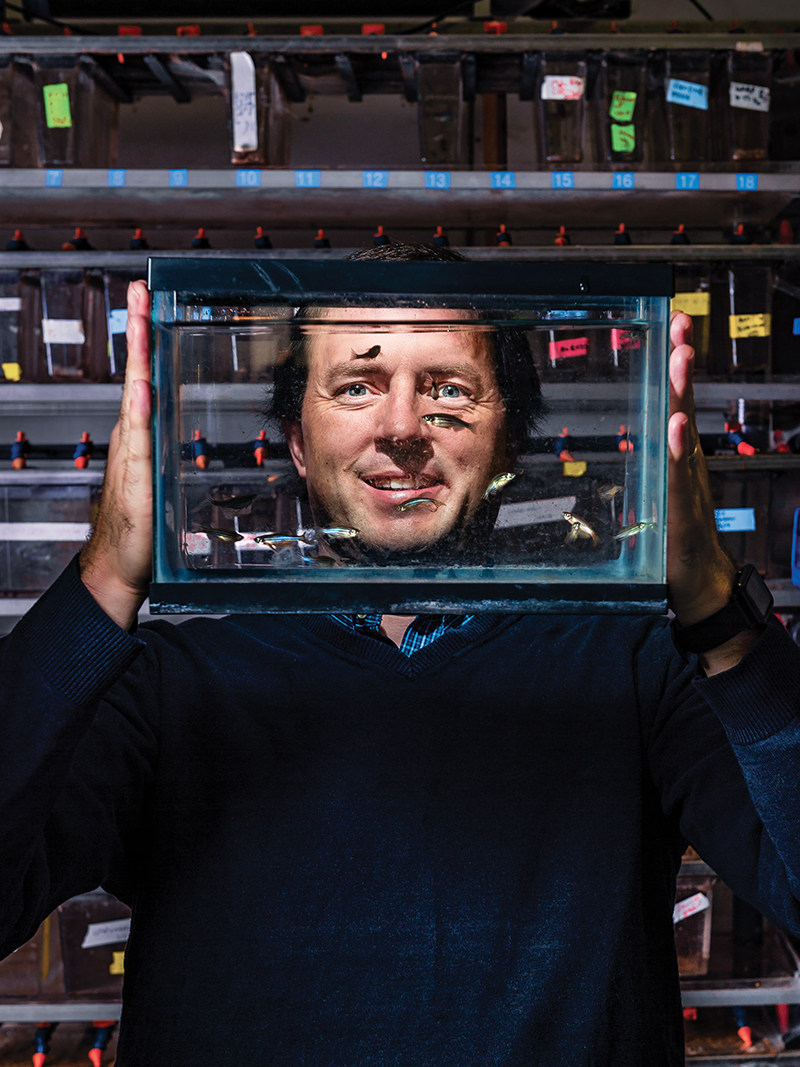

We humans have some limited powers of regeneration. After all, if we cut ourselves, we can regenerate skin to cover the wound. But that’s nothing like what zebrafish can do. “They can regenerate almost anything that doesn’t kill them,” says Jason Meyers, associate professor of biology and neuroscience and chair of scientific perspectives core. “They’ll regenerate their fins, their hearts, their retinas … if a needle is put down into their brain, they’ll even regenerate that.”
The bigger question, of course, is just how do they do it? Meyers has been using CRISPR to examine the genetics behind regeneration in zebrafish and what lessons it might hold for humans and other species. In many ways, zebrafish make the perfect species to investigate. Their body plan and nervous system are very similar to humans, they are transparent, and they are fast growing, so experiments can be carried out quickly. “It’s great for working with undergraduates, because you start an experiment on Monday and you can see exactly what happened by Thursday,” Meyers says.
Avery went to medical school but got frustrated at his inability to treat many diseases. That’s often how research starts — there is usually a big question we are trying to solve that drives us to study one of those unexplained mysteries. In his case, it made him realize he could tackle the really big questions in science.
— Jason Meyers, associate professor of biology and neuroscience and chair of scientific perspectives core
Scientists have also developed transgenic lines of zebrafish, which have been genetically modified to produce fluorescent-colored proteins in red and green that enable biologists to track exactly what genes do. Using such tools, Meyers has looked at how cells communicate with one another to spur the regeneration process. “How does a cell know that it’s been injured and needs to regenerate?” Meyers asks. By changing genes with CRISPR and tracking the different proteins, he has focused on a sequence of chemical interactions in the cells called the WNT signaling pathway, which seems to be important in telling the fish that it needs to regenerate lost structures such as retina or an earlike structure called the lateral line.
“If we turn the pathway up, we can overdrive the regenerative response and produce more than would normally be produced,” he says. With the help of students, he is now looking at other chemical pathways in the fish that might interact with the WNT pathway to complement the reaction. “Students design the pieces of RNA to inject, screen fish for mutations, do DNA sequencing to understand what the specific mutation did, and make movies tracking the development and regeneration of these systems,” Meyers says.
In addition to understanding how cells signal the regenerative process, Meyers is also examining whether the regeneration of organs is a recapitulation of the development from an embryo, or a completely different process. So far, the answer seems to be somewhere between the two.
Answering questions like this can help us better understand how stem cells might spur regeneration of organs, limbs, or brain cells in humans. “By pointing at particular pathways that seem to coordinate regeneration in zebrafish, we can find good places where we should start looking in mammalian systems,” Meyers says. “We are trying to lay down a foundation of knowledge about when and where regeneration happens, so we can better understand our own limitations — and possibly how to overcome them.”
For the past 13 years, Colgate’s biology department has held the Oswald T. Avery Lecture, delivered annually by an alumna or alumnus who has made significant contributions to the field.